Mutational hotspots are the result of a combination of evolutionary forces, including mutation rate heterogeneity. They have the ability to predict evolutionary outcomes and make evolution repeatable. However, hotspots are frequently difficult to locate.
Researchers discovered evolutionary hotspots in DNA where mutations are more likely to occur. According to the study’s authors, these findings will aid in the future prediction of the evolution of bacteria and viruses over time, which may aid in vaccine design and a better understanding of antibiotic resistance.
Tangles in unwound DNA can cause mutational hotspots in bacterial genomes, according to a new study from the University of Bath’s Milner Centre for Evolution. According to the study’s authors, these findings will aid in the future prediction of the evolution of bacteria and viruses over time, which may aid in vaccine design and a better understanding of antibiotic resistance.
Mutational hotspots have the potential to determine evolutionary outcomes and make evolution repeatable. Hotspots are the result of multiple evolutionary forces, including mutation rate heterogeneity, which is often difficult to identify.
Dr Tiffany Taylor
While most evolution is shaped by natural selection, in which only those individuals who are adapted to their environment survive and pass on their genes, a new study published in Nature Communications shows that tangles in DNA strands also influence evolution.
“Mutational hotspots have the potential to determine evolutionary outcomes and make evolution repeatable.” “Hotspots are the result of multiple evolutionary forces, including mutation rate heterogeneity, which is often difficult to identify,” the researchers wrote. “We show in this paper that a near-deterministic genetic hotspot can be built and broken by a small number of silent mutations.”
To understand the differences observed, the researchers compared the DNA sequences of the two strains. They discovered a region in the SBW25 strain, which mutated predictably, where the DNA strand looped back on itself, forming a hairpin-shaped tangle.
The evolution of two strains of the soil bacteria Pseudomonas fluorescens was studied by a team of scientists led by the University of Bath in collaboration with the University of Birmingham (SBW25 and Pf0-1).
When the scientists removed a gene that allows the bacteria to swim, both strains quickly evolved the ability to swim again, albeit in very different ways. To regain mobility, one of the strains (called SBW25) always mutated the same part of a specific gene. The other strain (dubbed Pf0-1), on the other hand, mutated in different places in different genes each time the scientists repeated the experiment.

They compared the DNA sequences of the two strains to figure out why one evolved predictably and the other was unpredictable. They discovered a region in the SBW25 strain, which mutated predictably, where the DNA strand looped back on itself, forming a hairpin-shaped tangle. These tangles can disrupt the cell machinery known as DNA polymerase, which copies the gene during cell division, increasing the likelihood of mutations.
When the team removed the hairpin structure with six silent mutations (without changing the sequence of the protein produced), the mutational hotspot was eliminated, and the bacteria began evolving in a much broader range of ways to regain its swimming ability.
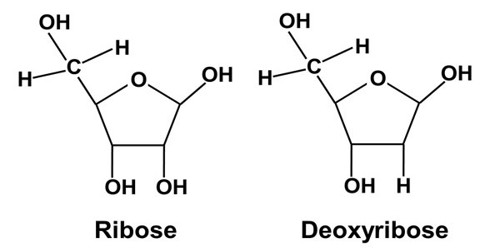
“DNA normally forms a double helix structure,” said Dr Tiffany Taylor of the Milner Centre for Evolution, “but when the DNA is copied, the strands are briefly separated. “We discovered hotspots in the DNA where the sequence causes the separated strands of DNA to twist back on themselves, similar to when you pull apart the strands of a rope, resulting in a tangle. When the DNA polymerase enzyme runs along the strand to copy the gene, it may come into contact with the tangle and skip, resulting in a mutation. We were able to create or remove mutational hotspots in the genome by changing the sequence to cause or prevent the hairpin tangle, according to our experiments. This demonstrates that, while natural selection remains the most important factor in evolution, there are other factors at work as well. If we knew where the potential mutational hotspots in bacteria or viruses were, we could predict how these microbes might mutate under selective pressure.”
Mutational hotspots have already been discovered in cancer cells, and the researchers intend to look for them in a variety of bacterial species, including pathogens. This knowledge can help scientists better understand how bacteria and viruses evolve, which can aid in the development of vaccines against new disease variants. It can also help predict how microbes will develop antibiotic resistance.
Dr. James Horton, a recent PhD graduate of the Milner Centre for Evolution, stated: “This, like many other exciting discoveries, was discovered by chance. Because the mutations we were looking at were so-called silent because they didn’t change the resulting protein sequence, we didn’t think they were particularly significant at first. Our findings, however, fundamentally challenge our understanding of the role of silent mutations in adaptation.”