Genetic disorders endanger farm animals’ welfare and have an impact on their production and management. One method of lowering this risk is to map the genes responsible for various syndromes. Until now, however, this has been a reactive process, with farmers notifying veterinarians or breeding companies when a genetic disorder is suspected, and researchers attempting to confirm the disorder by gathering information on the phenotypes and genotypes of selected individuals in order to identify the cause.
A new study published in the prestigious scientific journal Nature Genetics by Massey University and Livestock Improvement Corporation (LIC) researchers has now turned this process on its head, leveraging ever-growing genomic sequence data to proactively investigate animals’ differences in production traits caused by genome sequence variants. After identifying these outlier animals, further investigation revealed effects that had previously gone unnoticed.
Researchers leveraging ever-growing genomic sequence data to proactively investigate animals’ differences in production traits resulting from genome sequence variants.
The paper’s lead author, Edwardo Reynolds, a Ph.D. student at Massey’s AL Rae Centre of Genetics and Breeding in Ruakura, says, “These are fascinating discoveries. It demonstrates that we can identify new genetic disorders proactively and that there is a viable way to improve the health and productivity of New Zealand dairy animals.”
Matt Littlejohn, LIC’s research leader and Massey Professor in Animal Genetics, oversaw the study and says, “We were taken aback by the number and magnitude of the effects. Edwardo’s publication of this work in one of the world’s best biology journals is also a significant accomplishment for a scientist in his early career.”
“The research demonstrates the value of academia and industry working together and the benefits that can be developed for farmers when pure and applied sciences are used to tackle real-world problems,” says Professor Dorian Garrick, co-supervisor of Mr. Reynolds and chief scientist at the AL Rae Centre.
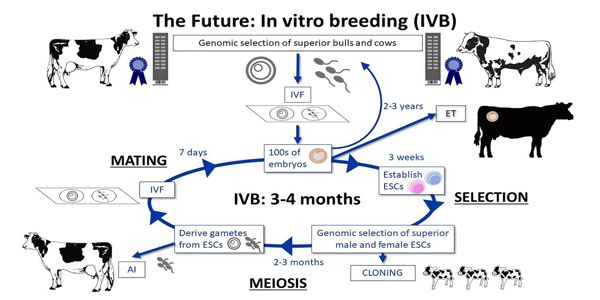
The process of determining the location of genes on each chromosome is known as genome mapping. The maps produced are comparable to the maps we use to navigate streets. A genetic map is a diagram that depicts genes and their locations on chromosomes. Genetic maps provide a big picture (similar to an interstate highway map) and make use of genetic markers (similar to landmarks). A genetic marker is a gene or sequence on a chromosome that demonstrates a genetic link to an interesting trait.
The researchers discovered six recessive variants with effects ranging from mild (slight loss of body condition with limited other consequences) to severe (approximately 25 percent reduction in body weight and increased early-life mortality).
Aside from the bodyweight effects that led to the discoveries, many other effects were discovered, such as lower milk production (approximately 1000 liters of milk per lactation for the biggest effect variant), lower milk solids (75kg less per lactation for the biggest effect variant), smaller stature, smaller chest circumference, and other anatomical changes.
According to Professor Littlejohn, knowledge of these variants can now be used to help manage the variants’ frequencies through genetic testing.
The outline is provided by genetic maps, while the details are provided by physical maps. It’s easy to see why both types of genome-mapping techniques are important for revealing the big picture. The information obtained from each technique is combined to study the genome. Genomic mapping is used in research with a variety of model organisms. Genome mapping is still an ongoing process, and more advances are expected as more advanced techniques are developed. Genome mapping is similar to completing a complicated puzzle using every piece of available data.