Proteins are the fundamental components of all living organisms. They are essential for the construction, function, and control of the body’s tissues and organs, and they conduct the majority of their activity in cells. From enzymes that carry out chemical processes to messengers that transfer messages between cells, a great deal of study is done on how these proteins are generated and what they accomplish.
The Nobel Prize in Chemistry was awarded to Aaron Ciechanover, Avram Hershko, and Irwin Rose in 2004 for a separate but equally vital function of protein machinery: how organisms break down proteins once they have completed their task.
Protein degradation is a well-planned procedure. Proteins are labeled with ubiquitin, a chemical marker for disposal, and then fed into proteasomes, a type of cellular paper shredder that breaks up proteins into minute bits.
Ubiquitination, or the act of tagging proteins with ubiquitin, is involved in a variety of biological functions, such as cell division, DNA repair, and immunological responses. Proteins are made up of hundreds or thousands of smaller components known as amino acids that are linked in lengthy chains.
Researchers from the University of Chicago employed powerful electron microscopes to explore further into the process of protein breakdown in a new study published in Nature on November 17, 2021.
They discovered the structure of a crucial enzyme that aids in the mediation of ubiquitination in yeast, which is part of a biological mechanism known as the N-degron pathway that may determine the rate of breakdown for up to 80% of comparable proteins in humans.
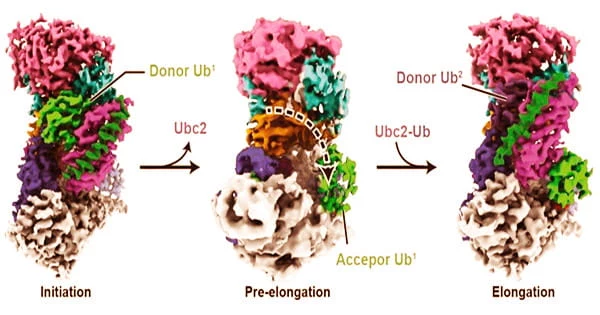
Because malfunctions in this system can result in a buildup of damaged or misfolded proteins, which is linked to the aging process, neurodegeneration, and several uncommon autosomal recessive illnesses, a better knowledge of it could lead to new therapies.
Until this study, we didn’t know that much about how ubiquitin polymers are structurally formed. Now we are starting to get an idea of how it’s first installed onto the protein substrate, and then how the polymers are formed in a linkage-specific manner. This is a milestone in terms of understanding polyubiquitination at a near-atomic level.
Minglei Zhao
Proteins are involved in practically every aspect of cellular life, including cell structure and organization, product production and waste disposal, and routine maintenance. They also receive signals from outside the cell and activate a reaction within the cell.
Minglei Zhao, PhD, Assistant Professor of Biochemistry and Molecular Biology, and his colleagues investigated Ubr1, an E3 ligase enzyme that aids in the joining of bigger molecules. Ubr1 aids in the ubiquitination process in baker’s yeast by attaching ubiquitin to proteins and elongating it into a polymer chain of molecules.
Polymers, which are more frequently associated with manufactured materials like plastics, also arise naturally when big molecules (in this case, ubiquitin) are linked in repeating subunits.
“Until this study, we didn’t know that much about how ubiquitin polymers are structurally formed,” Zhao said. “Now we are starting to get an idea of how it’s first installed onto the protein substrate, and then how the polymers are formed in a linkage-specific manner. This is a milestone in terms of understanding polyubiquitination at a near-atomic level.”
Zhao and his colleagues employed chemical biology techniques to imitate the earliest phases of the ubiquitin-protein attachment pathway. They then used cryo-electron microscopy (cryo-EM), a Nobel Prize-winning technique, to capture the process. Cryo-EM involves flash-freezing protein solutions and then imaging individual molecules or subcellular structures with a powerful electron microscope.
Breakthroughs in technology and software about ten years ago resulted in microscopes and detectors that could capture molecular pictures with considerably better resolution. Cryo-EM methods, which allow researchers to take a picture that essentially freezes “live” activity of a biological process, were awarded the Nobel Prize in Chemistry in 2017 to Jacques Dubochet, Joachim Frank, and Richard Henderson.
Zhao’s team used cryo-EM to examine ubiquitination in more depth, thanks to a $10 million investment from the Biological Sciences Division in the Advanced Electron Microscope Facility.
They were able to characterize the structure of numerous intermediary enzyme complexes involved in the pathway, which will aid researchers exploring for new ways to target proteins with medications or intervene in a faulty protein breakdown process.
“Cryo-EM is exciting because after the data processing is done, a new structure pops out that you’ve never seen before,” Zhao said. “Now we can use what we’ve learned and repurpose the enzymes by introducing small molecules or mixture of peptides to degrade the proteins we want.”