According to a new study, antibiotic resistance in E. coli has been steadily increasing since the early 2000s, despite efforts to control it. Researchers successfully tracked the spread of antibiotic resistant genes and demonstrated that these genes are being transferred between E. coli strains in the largest genomic survey of E. coli to date, which took more than 16 years in Norway.
Researchers from the Wellcome Sanger Institute and the University of Oslo studied multidrug resistance in Norway and compared it to a previous study conducted in the United Kingdom. They discovered that resistant strains appeared around the same time but spread more quickly in the UK population.
The findings, published today in The Lancet Microbe, demonstrate the importance of tracking these resistant strains in the surveillance and control of drug-resistant E. coli, which is a major problem in hospitals where it can cause severe infection and mortality. Furthermore, understanding how these genes are transferred between strains and what caused them to develop drug resistance can aid in the prevention of the spread of antibiotic resistance strains.
Researchers have tracked multidrug resistance in Norway and compared this to a previous study from the UK. They found that resistant strains developed around the same time, but increased more rapidly in the UK population.
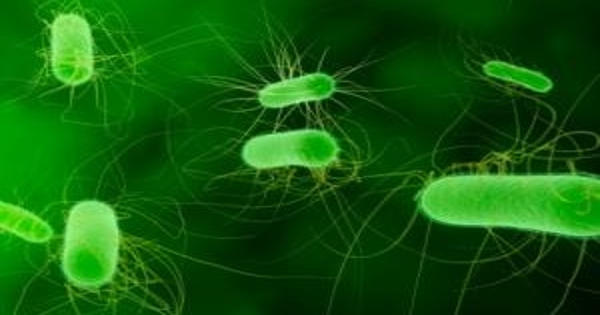
The bacterium Escherichia coli is a common cause of bloodstream infections around the world, and its prevalence appears to be increasing over the last decade. E. coli is commonly found in the gut, where it is harmless, but if it enters the bloodstream due to a weakened immune system, it can cause severe and life-threatening infections. Multi-drug resistance (MDR) has become a common feature of such infections, posing an additional challenge for health care providers, and in a worrying number of cases, treatment options are becoming limited.
Researchers from the Wellcome Sanger Institute and the University of Oslo processed a nationwide catalog of samples from more than 3,200 patients to track antibiotic resistance over 16 years in the largest study of its kind and only the second systematic longitudinal genomic study of bacteremia E. coli. They tracked the emergence of drug resistance using the power of large-scale DNA sequencing and compared it to a similar study conducted in the UK.
The researchers discovered that MDR began to increase and appear in more strains in the early 2000s as a result of antibiotic pressure and that multiple MDR E. coli strains are now present in Norway. However, despite similar antibiotic-use policies, MDR E. coli appears to be more prevalent in the UK. The UK population, on the other hand, is significantly larger than Norway’s, which could explain some of the differences. More research is needed to allow for more precise comparisons and to identify the specific factors that cause rapid spread in some locations versus others.
MDR is relatively uncommon in bacteria. However, this new study discovered that lineages previously thought to lack MDR have acquired drug-resistance genes, demonstrating E. coli’s increased ability to share MDR genes that move horizontally between strains.
According to Professor Jukka Corander, co-author and Associate Faculty member at the Wellcome Sanger Institute: “Because of the large number of samples from the Norwegian population and the level of genomic detail on the bacteria strains, we were able to draw far-reaching conclusions that were previously unthinkable. This study demonstrates the power of systematic national surveillance of resistant organisms that collects and makes data available for in-depth analysis. Without these, it would have been impossible to approach the study’s central research questions and find answers to them.”
The researchers hope to conduct similar research in the UK to build on previous studies and gain a full data set of 16 years in the UK in order to more closely track MDR-resistant E. coli.
Dr. Rebecca Gladstone, the study’s lead author, and a bioinformatician at the University of Oslo in Norway stated: “The ability to estimate the expansion timelines of E. coli MDR clones and identify multiple occasions of novel resistance gene acquisition is especially exciting because it is the first time this has been possible. Understanding and tracking the movement of these drug resistance genes and the strains that carry them is critical for controlling the spread of drug-resistant bacteria, which is a major problem in the healthcare industry.”
Professor Julian Parkhill, co-author, and Professor at the University of Cambridge’s Department of Veterinary Medicine stated: “Long-term studies like this one provide a thorough understanding of the complex epidemiology that underpins bloodstream infections. Further research to detail the factors determining the success of emerging pathogenic clones of these bacteria would be the next step, in order to help find a way to control and possibly reduce the spread of multidrug resistance.”