Researchers frequently employ two-photon microscopy to generate high-resolution, 3D images of tissues like the brain, which requires directing a high-intensity laser onto the material to cause fluorescence excitation. Scanning deep into the brain, on the other hand, can be challenging because light bounces off of tissues as it travels deeper, resulting in hazy pictures. Two-photon imaging takes time since it involves scanning individual pixels one at a time.
A group of researchers from MIT and Harvard University has created a modified version of two-photon imaging that can scan deeper inside tissue and much more swiftly than previously feasible. According to the researchers, this type of imaging might allow scientists to get high-resolution pictures of features such as blood arteries and individual neurons within the brain more quickly.
“By modifying the laser beam coming into the tissue, we showed that we can go deeper and we can do finer imaging than the previous techniques,” says Murat Yildirim, an MIT research scientist and one of the authors of the new study.
The paper’s lead authors are MIT graduate student Cheng Zheng and former postdoc Jong Kang Park, and it was published on 7th July (2021) in Science Advances. The paper’s senior author is Dushan N. Wadduwage, a former MIT postdoc who is currently a John Harvard Distinguished Science Fellow in Imaging at Harvard University’s Center for Advanced Imaging. Josiah Boivin, an MIT postdoc; Yi Xue, a former MIT graduate student; Mriganka Sur, MIT’s Newton Professor of Neuroscience; and Peter So, an MIT professor of mechanical and biological engineering, are among the other writers.
Deep imaging:
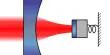
Two-photon microscopy works by focusing a powerful beam of near-infrared light onto a single spot within a sample, causing the simultaneous absorption of two photons at the focal point, where the intensity is greatest. This low-energy, long-wavelength light may penetrate deeper into tissue without causing damage, allowing imaging under the surface.
Two-photon excitation, on the other hand, produces pictures through fluorescence, and the fluorescent signal is visible. The fluorescent light scatters more as it penetrates deeper into tissue samples, resulting in a hazy picture. It takes a long time to image several layers of tissue. The procedure can be sped up by using wide-field imaging, which illuminates a whole plane of tissue at once, but the resolution isn’t as good as point-by-point scanning.
The MIT researchers intended to devise a method for imaging a large tissue sample at once while preserving the high resolution of point-by-point scanning. To do so, they devised a method of manipulating the light that they beam onto the sample. They employ a type of wide-field microscopy in which they shine a plane of light onto the tissue but change the amplitude of the light so that each pixel can turn on and off at various times. This predesigned pattern may be recognized in the light dispersed by the tissue since certain pixels are lighted up while neighboring pixels remain dark.
“We can turn each pixel on or off by this kind of modulation,” Zheng says. “By turning off part of the spots, we create space around each pixel, allowing us to see what’s going on in each individual place.”
After obtaining the raw pictures, the researchers recreate each pixel using a computer technique that they developed.
“We control the shape of the light and we get the response from the tissue. From these responses, we try to resolve what kind of scattering the tissue has. As we do the reconstructions from our raw images, we can get a lot of information that you cannot see in the raw images,” Yildirim says.
The researchers demonstrated that they could scan 200 microns deep into slices of muscle and kidney tissue, as well as 300 microns deep into the brains of mice, using this approach. According to Yildirim, this is roughly twice as deep as could be achieved without the use of programmed stimulation and computer reconstruction. In addition, the method may produce pictures 100 to 1,000 times quicker than traditional two-photon microscopy.
Brain structure:
Researchers should be able to get high-resolution pictures of neurons in the brain, as well as other structures like blood arteries, more quickly using this sort of imaging. According to Yildirim, imaging blood arteries in the brains of mice might be particularly beneficial for understanding more about how neurodegenerative disorders like Alzheimer’s impact blood flow.
“All blood flow or morphological investigations of blood artery structures use two-photon or three-photon point scanning methods, which are slow,” he says. “We can really do high-speed volumetric imaging of blood flow and blood vessel anatomy using this technology in order to understand variations in blood flow.”
By using voltage-sensitive fluorescent dyes or fluorescent calcium probes that light up when neurons are stimulated, the method might potentially be used to measure neural activity. It may also be used to examine other forms of tissue, such as cancers, where it could be utilized to assist detect the tumor’s boundaries.